10 Regression
10.1 Install Packages
install.packages("Metrics")10.3 LINKS
Nice tutorial in R —>
Beginners Guide to Regression Analysis and Plot Interpretations
There’s an error in the above webpage according to the definition of \(R^2\), which is:
\[R^2 = 1 - \frac{RSS}{TSS} = \frac{ESS}{TSS}\]
10.4 Introduction
The aim of linear regression is to model a continuous variable Y as a mathematical function of one or more X variable(s), so that we can use this regression model to predict the Y when only the X is known. This mathematical equation can be generalized as follows:
\[Y = \beta_{1} + \beta_{2}X + \epsilon\]
where, \(\beta_{1}\) is the intercept and \(\beta_{2}\) is the slope.
Collectively, they are called regression coefficients. \(\epsilon\) is the error term, the part of Y the regression model is unable to explain.
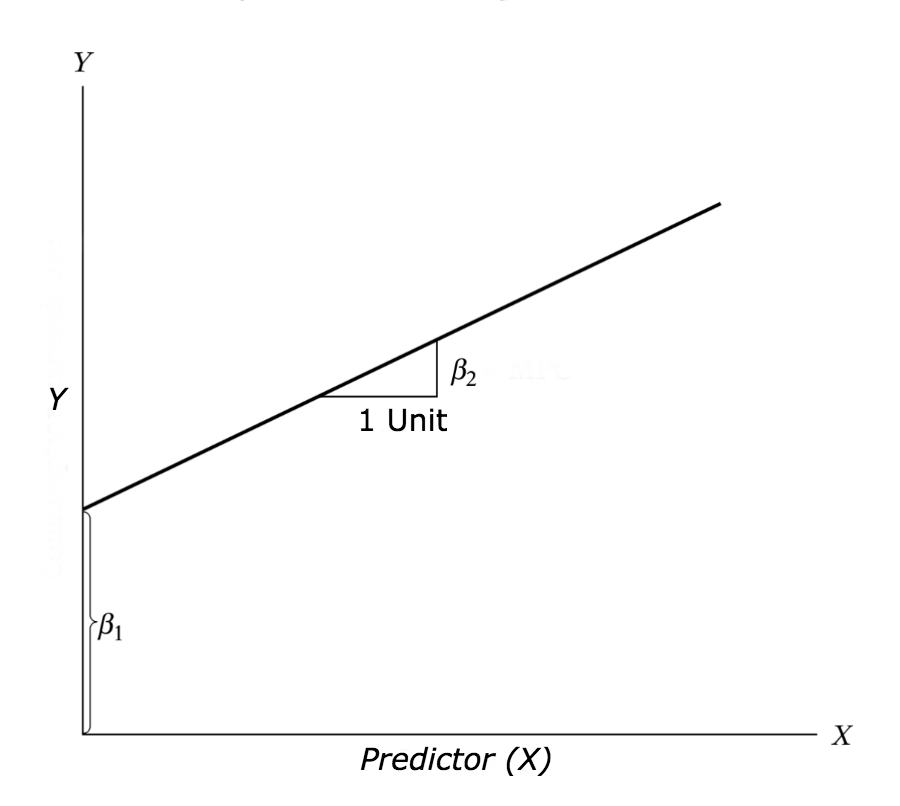
Figure 10.1: Linear regression explained.
10.5 Import Climatic Data [ \(T_{max}\), Rain ]
Tmax <- read_excel("../datasets/interchange/Tmax.xlsx", sheet = "measurements", na = "NA")
Rain <- read_excel("../datasets/interchange/Rainfall.xlsx", sheet = "measurements", na = "NA")10.6 Scatter Plot
scatter.smooth( x=Tmax$MODENAURB, y=Tmax$SAGATABOAGRO,
main="SAGATABOAGRO ~ MODENAURB")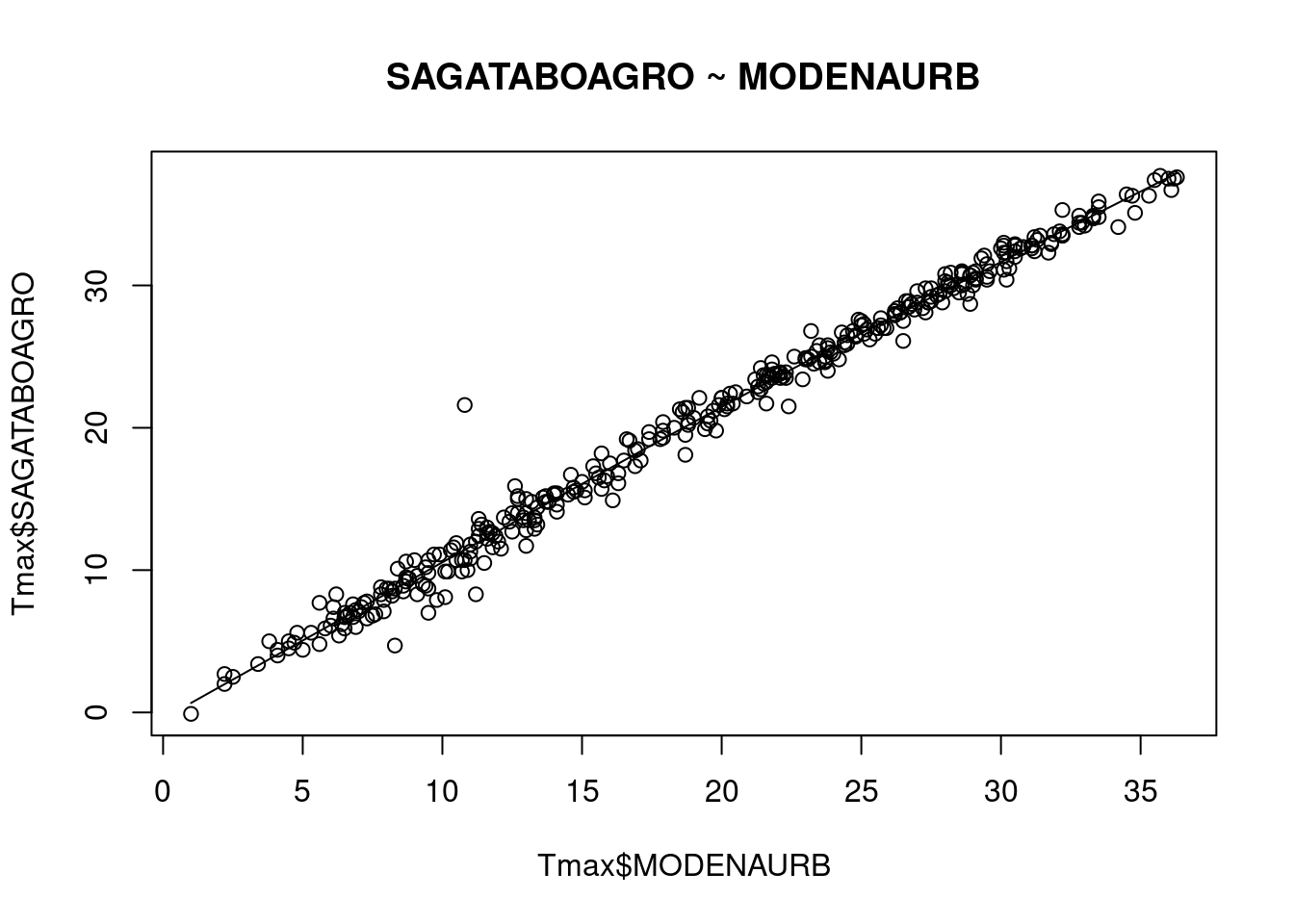
10.7 BoxPlot – Check for outliers
Generally, any datapoint that lies outside the 1.5 * interquartile-range (1.5 * IQR) is considered an outlier, where, IQR is calculated as the distance between the 25th percentile and 75th percentile values for that variable.
par(mfrow=c(1, 2)) # divide graph area in 2 columns
boxplot(Tmax$MODENAURB, main="MODENAURB",
sub=paste("Outlier rows: ", boxplot.stats(Tmax$MODENAURB)$out))
boxplot(Tmax$SAGATABOAGRO, main="SAGATABOAGRO",
sub=paste("Outlier rows: ", boxplot.stats(Tmax$SAGATABOAGRO)$out))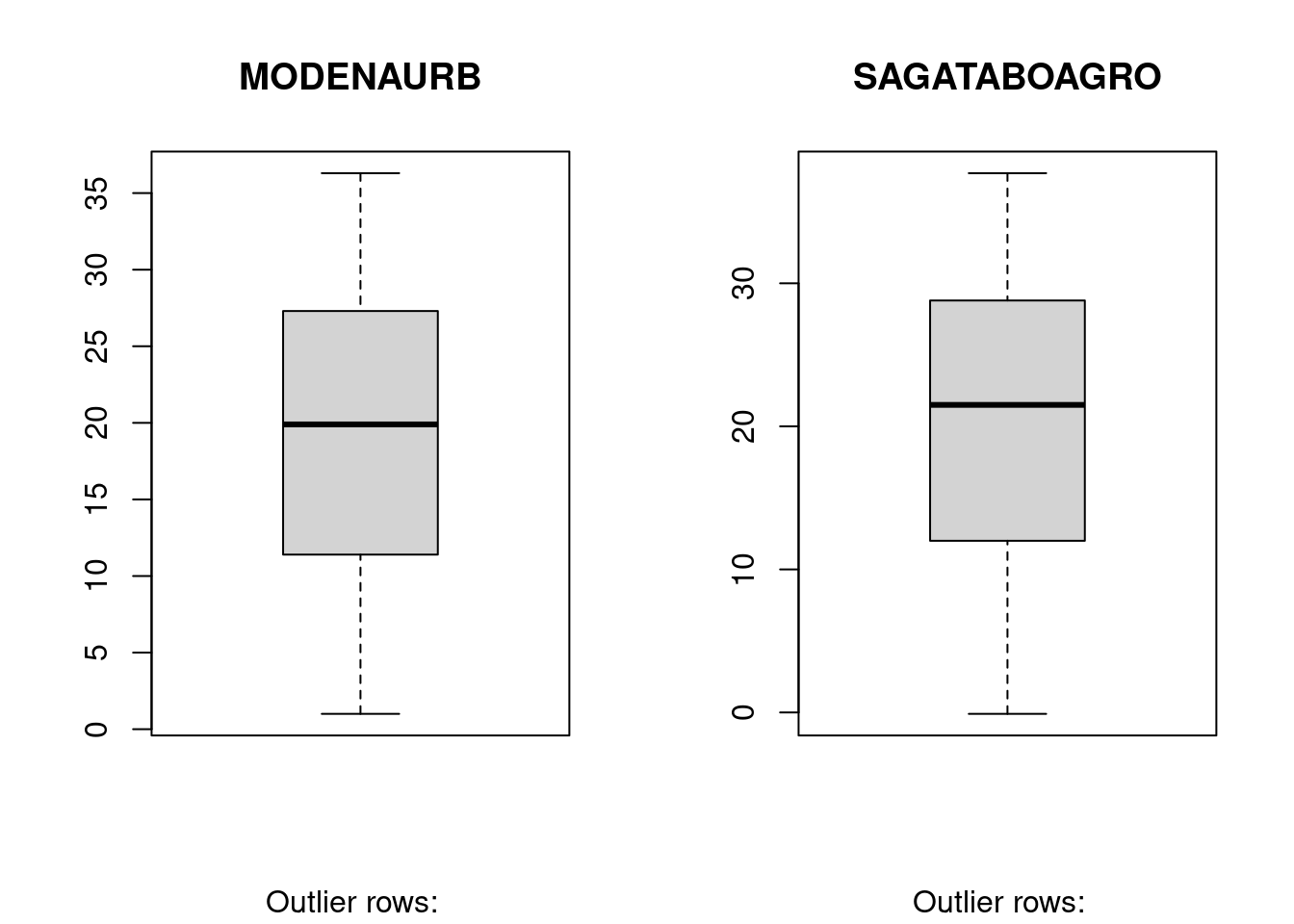
10.8 Density plot – Check if the response variable is close to normality
library(e1071)
par(mfrow=c(1, 2)) # divide graph area in 2 columns
plot(density(Tmax$MODENAURB),
main="Density Plot: MODENAURB", ylab="Frequency",
sub=paste("Skewness:", round(e1071::skewness(Tmax$MODENAURB), 2)))
polygon(density(Tmax$MODENAURB), col="red")
plot(density(Tmax$SAGATABOAGRO),
main="Density Plot: SAGATABOAGRO", ylab="Frequency",
sub=paste("Skewness:", round(e1071::skewness(Tmax$SAGATABOAGRO), 2)))
polygon(density(Tmax$SAGATABOAGRO), col="red")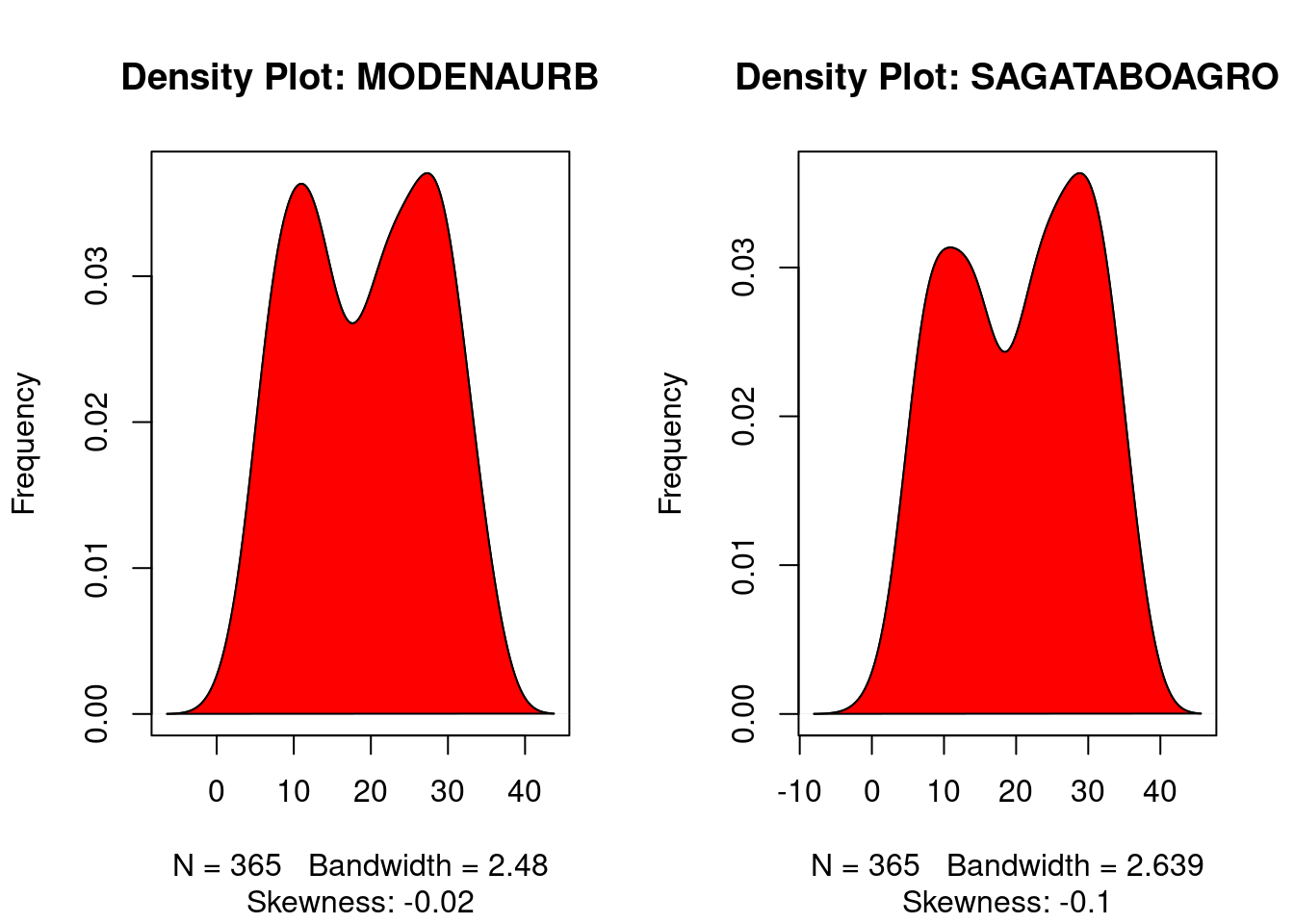
10.9 Build linear regression model on full data
linearMod <- lm(SAGATABOAGRO ~ MODENAURB, data=Tmax)
print(linearMod)
#>
#> Call:
#> lm(formula = SAGATABOAGRO ~ MODENAURB, data = Tmax)
#>
#> Coefficients:
#> (Intercept) MODENAURB
#> 0.03945 1.05787
summary(linearMod)
#>
#> Call:
#> lm(formula = SAGATABOAGRO ~ MODENAURB, data = Tmax)
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -4.1198 -0.4477 0.0439 0.4954 10.1355
#>
#> Coefficients:
#> Estimate Std. Error t value Pr(>|t|)
#> (Intercept) 0.039447 0.126553 0.312 0.755
#> MODENAURB 1.057875 0.005932 178.326 <2e-16 ***
#> ---
#> Signif. codes:
#> 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
#>
#> Residual standard error: 1.015 on 363 degrees of freedom
#> Multiple R-squared: 0.9887, Adjusted R-squared: 0.9887
#> F-statistic: 3.18e+04 on 1 and 363 DF, p-value: < 2.2e-16
linearMod$coefficients
#> (Intercept) MODENAURB
#> 0.03944719 1.05787488That is:
\(Y = \beta_{1} + \beta_{2}X + \epsilon\)
\(SAGATABOAGRO = Intercept + (\beta ∗ MODENAURB)\)
\(SAGATABOAGRO = +0.03945 + 1.05787* MODENAURB\)
- \(y_{i}\) observations;
- \(\overline{y}\) the average value;
- \(\hat{y_{i}}\) estimation from linear regression.
yi = Tmax$SAGATABOAGRO
y_ = mean(yi)
y_hat = +0.03945 + 1.05787*Tmax$MODENAURB10.9.1 Graphical visualization:
ggplot(Tmax, aes(x=MODENAURB, y=SAGATABOAGRO)) +
geom_point(shape=18) +
geom_smooth(method = "lm", se = FALSE, color="red") +
annotate("segment", x = 0, xend = max(Tmax$MODENAURB),
y = mean(Tmax$SAGATABOAGRO),
yend = mean(Tmax$SAGATABOAGRO), colour = "blue") +
annotate("text", x = 0, y = 30, label = "yi : dati osservati",
colour="black", hjust=0, vjust=0) +
annotate("text", x = 0, y = 28, label = "y_hat : dati stimati dal modello",
colour="red", hjust=0, vjust=0) +
annotate("text", x = 0, y = 26, label = "y_ : valore medio",
colour="blue", hjust=0, vjust=0)
#> `geom_smooth()` using formula = 'y ~ x'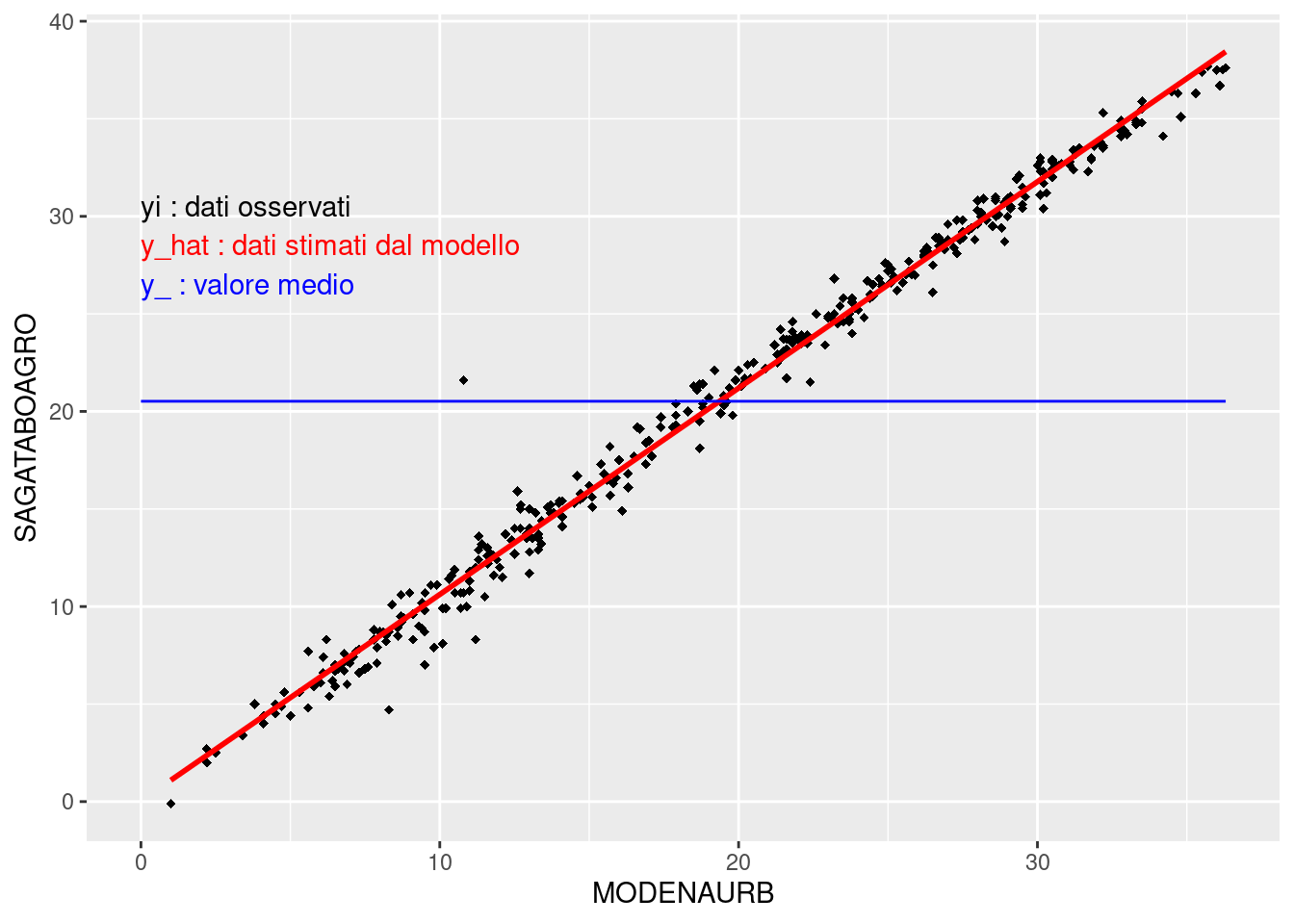
10.9.2 Deviance explained by the regression model (Explained Sum of Squares):
\[ESS = \sum_{i=1}^{n}\left( \hat{y_{i}}-\bar{y}\right)^{2}\]
10.9.3 Total Deviance (Total Sum of Squares)
\[TSS = \sum_{i=1}^{n}\left( y_{i}-\bar{y}\right)^{2}\]
10.9.4 Coefficient of determination
\[R^{2}=\frac{ESS}{TSS}\]
R2 = ESS / TSS
print(R2)
#> [1] 0.988704710.10 Linear regression model to infill missing data
10.10.1 Selezione della variabile dipendente o target da ricostruire:
yi = Tmax$SAGATABOAGRO # copia di SAGATABOAGRO da manipolare!
mean(yi)
#> [1] 20.5216410.10.2 Rimozione di alcuni dati misurati, per simulare la presenza di dati mancanti:
NA_POS = c(4,7,22,35,48,56,78,89,94,102,134,157,187,232,259,299,305,360)
MEAS_DATA = yi[NA_POS]
print( paste("No. of removed measurements: ",length(NA_POS)) )
#> [1] "No. of removed measurements: 18"
MEAS_DATA
#> [1] 7.0 9.9 7.8 8.8 12.7 11.1 16.3 9.0 14.8 23.4 31.1
#> [12] 20.7 32.8 26.1 30.0 12.4 14.8 5.910.10.2.1 NA_POS
sample(1:365,20,replace=FALSE)
#> [1] 81 166 30 339 53 289 8 338 147 268 274 349 116 173
#> [15] 180 94 89 247 151 7510.10.4 Ricostruzione dei dati mancati: calcolo della media mobile
y_mm = (yi[NA_POS-1] + yi[NA_POS+1] ) / 2
y_mm
#> [1] 12.40 9.45 12.00 12.50 11.65 14.45 13.40 15.20 19.60
#> [10] 24.10 27.65 24.95 31.20 26.90 28.75 13.50 16.75 5.25
yi[NA_POS[1]]
#> [1] NA10.10.5 Valori misurati rimossi, negli stessi giorni:
MEAS_DATA
#> [1] 7.0 9.9 7.8 8.8 12.7 11.1 16.3 9.0 14.8 23.4 31.1
#> [12] 20.7 32.8 26.1 30.0 12.4 14.8 5.910.10.6 Ricostruzione dei dati mancati: regression lineare
10.10.6.1 Matrice X delle variabili indipendenti o covariate usate per ricostruire i dati mancanti
# Selezione delle stazioni di MODENAURB (pos=2) e CORREGGIOAGRO (pos=6)
X = Tmax[,c(2,6)]
summary(X)
#> MODENAURB CORREGGIOAGRO
#> Min. : 1.00 Min. :-0.20
#> 1st Qu.:11.40 1st Qu.:11.50
#> Median :19.90 Median :21.00
#> Mean :19.36 Mean :20.15
#> 3rd Qu.:27.30 3rd Qu.:28.50
#> Max. :36.30 Max. :37.1010.10.6.2 Costruzione del modello di regressione tra la stazione da ricostruire e le altre disponibili:
yi[NA_POS] # questi sono i valori mancanti da ricostruire!
#> [1] NA NA NA NA NA NA NA NA NA NA NA NA NA NA NA NA NA NA
# build linear regression model using 1 covariate
lm1 <- lm(yi ~ X$MODENAURB)
print(lm1)
#>
#> Call:
#> lm(formula = yi ~ X$MODENAURB)
#>
#> Coefficients:
#> (Intercept) X$MODENAURB
#> 0.08073 1.05715
# build linear regression model using 2 covariate
lm2 <- lm(yi ~ X$MODENAURB + X$CORREGGIOAGRO)
print(lm2)
#>
#> Call:
#> lm(formula = yi ~ X$MODENAURB + X$CORREGGIOAGRO)
#>
#> Coefficients:
#> (Intercept) X$MODENAURB X$CORREGGIOAGRO
#> 0.1140 0.3612 0.6662
y_hat_1 = 0.06158 + 1.05783*X$MODENAURB[NA_POS]
y_hat_2 = 0.07883 + 0.35696*X$MODENAURB[NA_POS] + 0.67155*X$CORREGGIOAGRO[NA_POS]
#y_hat_2_bis = beta(1) + beta(2)*X$MODENAURB[NA_POS] + beta(3)*X$CORREGGIOAGRO[NA_POS]
beta = coefficients(lm1)
beta
#> (Intercept) X$MODENAURB
#> 0.08073207 1.05714879
y_hat_1 = beta[1] + beta[2]*X$MODENAURB[NA_POS]
y_hat_1
#> [1] 10.123646 10.757935 7.797918 8.326493 13.295092
#> [6] 10.546505 16.783683 9.912216 14.669385 24.289439
#> [11] 31.900911 20.166559 32.958060 28.095175 30.738047
#> [16] 12.660803 14.563671 6.952199
beta = coefficients(lm2)
beta
#> (Intercept) X$MODENAURB X$CORREGGIOAGRO
#> 0.1139654 0.3611984 0.6662450
y_hat_2 = beta[1] + beta[2] * X$MODENAURB[NA_POS] + beta[3] * X$CORREGGIOAGRO[NA_POS]
y_hat_2
#> [1] 8.209065 10.757641 7.880800 8.128024 12.957007
#> [6] 9.819283 16.480819 9.269442 15.158802 24.308663
#> [11] 31.373133 20.168385 32.800323 27.807586 29.976447
#> [16] 12.007419 15.322556 7.125470
cbind(y_,y_mm,MEAS_DATA,y_hat_1,y_hat_2)
#> y_ y_mm MEAS_DATA y_hat_1 y_hat_2
#> [1,] 20.73718 12.40 7.0 10.123646 8.209065
#> [2,] 20.73718 9.45 9.9 10.757935 10.757641
#> [3,] 20.73718 12.00 7.8 7.797918 7.880800
#> [4,] 20.73718 12.50 8.8 8.326493 8.128024
#> [5,] 20.73718 11.65 12.7 13.295092 12.957007
#> [6,] 20.73718 14.45 11.1 10.546505 9.819283
#> [7,] 20.73718 13.40 16.3 16.783683 16.480819
#> [8,] 20.73718 15.20 9.0 9.912216 9.269442
#> [9,] 20.73718 19.60 14.8 14.669385 15.158802
#> [10,] 20.73718 24.10 23.4 24.289439 24.308663
#> [11,] 20.73718 27.65 31.1 31.900911 31.373133
#> [12,] 20.73718 24.95 20.7 20.166559 20.168385
#> [13,] 20.73718 31.20 32.8 32.958060 32.800323
#> [14,] 20.73718 26.90 26.1 28.095175 27.807586
#> [15,] 20.73718 28.75 30.0 30.738047 29.976447
#> [16,] 20.73718 13.50 12.4 12.660803 12.007419
#> [17,] 20.73718 16.75 14.8 14.563671 15.322556
#> [18,] 20.73718 5.25 5.9 6.952199 7.12547010.10.7 Create two functions for error measurement:
Examples about how to create functions tailored to our needs.
10.10.9 Calculate MAE
mae(MEAS_DATA,y_)
#> [1] 8.783189
mae(MEAS_DATA,y_mm)
#> [1] 2.655556
mae(MEAS_DATA,y_hat_1)
#> [1] 0.7664819
mae(MEAS_DATA,y_hat_2)
#> [1] 0.5973194| y_ | MEAS_DATA | y_mm | y_hat_1 | y_hat_2 | |||
|---|---|---|---|---|---|---|---|
| 20.73718 | 7.0 | 12.40 | 5.40 | 10.123646 | 3.1236456 | 8.209065 | 1.2090649 |
| 20.73718 | 9.9 | 9.45 | -0.45 | 10.757935 | 0.8579349 | 10.757641 | 0.8576413 |
| 20.73718 | 7.8 | 12.00 | 4.20 | 7.797918 | -0.0020817 | 7.880800 | 0.0807999 |
| 20.73718 | 8.8 | 12.50 | 3.70 | 8.326493 | -0.4735073 | 8.128024 | -0.6719764 |
| 20.73718 | 12.7 | 11.65 | -1.05 | 13.295092 | 0.5950920 | 12.957007 | 0.2570074 |
| 20.73718 | 11.1 | 14.45 | 3.35 | 10.546505 | -0.5534949 | 9.819283 | -1.2807169 |
| 20.73718 | 16.3 | 13.40 | -2.90 | 16.783683 | 0.4836830 | 16.480819 | 0.1808194 |
| 20.73718 | 9.0 | 15.20 | 6.20 | 9.912216 | 0.9122159 | 9.269442 | 0.2694416 |
| 20.73718 | 14.8 | 19.60 | 4.80 | 14.669385 | -0.1306146 | 15.158802 | 0.3588022 |
| 20.73718 | 23.4 | 24.10 | 0.70 | 24.289440 | 0.8894395 | 24.308663 | 0.9086632 |
| 20.73718 | 31.1 | 27.65 | -3.45 | 31.900911 | 0.8009108 | 31.373133 | 0.2731329 |
| 20.73718 | 20.7 | 24.95 | 4.25 | 20.166559 | -0.5334408 | 20.168385 | -0.5316149 |
| 20.73718 | 32.8 | 31.20 | -1.60 | 32.958060 | 0.1580596 | 32.800323 | 0.0003233 |
| 20.73718 | 26.1 | 26.90 | 0.80 | 28.095175 | 1.9951751 | 27.807586 | 1.7075858 |
| 20.73718 | 30.0 | 28.75 | -1.25 | 30.738047 | 0.7380471 | 29.976447 | -0.0235528 |
| 20.73718 | 12.4 | 13.50 | 1.10 | 12.660803 | 0.2608027 | 12.007419 | -0.3925811 |
| 20.73718 | 14.8 | 16.75 | 1.95 | 14.563671 | -0.2363294 | 15.322556 | 0.5225558 |
| 20.73718 | 5.9 | 5.25 | -0.65 | 6.952199 | 1.0521992 | 7.125470 | 1.2254697 |
10.10.10 Use our function to measure the error
rmse(MEAS_DATA,y_)
#> [1] 9.61955310.10.11 Use a dedicated package to measure the error
Metrics::rmse(MEAS_DATA,y_)
#> [1] 9.619553